New algorithms identify measuring errors in single-cell sequencing
AI finds errors in RNA analysis
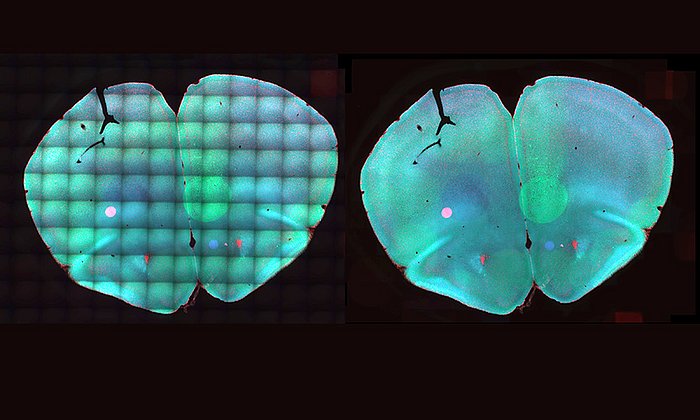
Being able to map all the cells in the human body and thereby improve the diagnosis, monitoring, and treatment of diseases — this is the vision behind the international Human Cell Atlas project. Such a reference database for the development of personalized medicine aims to allow healthy cells to be distinguished from diseased ones. This is made possible by single-cell RNA sequencing. With this method, it can be determined which genes play a role for the production of a cell. When a protein for cell assembly is generated, only certain segments of a person's DNA are read and translated into RNA which serves as a basis for protein biosynthesis.
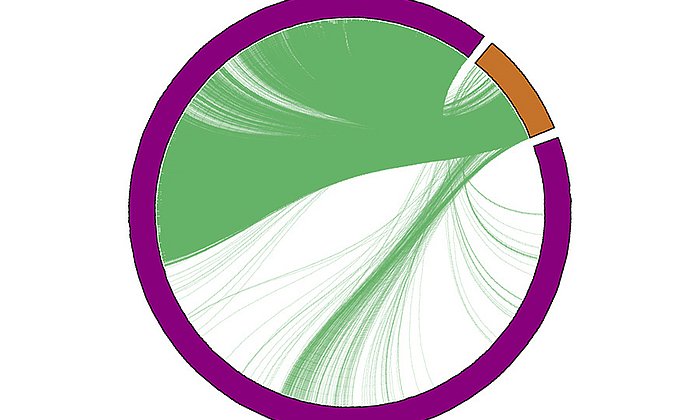
Extremely fine measurements are necessary for single-cell RNA sequencing. These are frequently distorted by the devices used, the environment, or the cell biology itself. Discrepancies in the measurements occur, for example, when the temperature of the measuring instrument has deviated even slightly or the processing time of the cells changes. Although several models exist for the correction of this so called batch effect, those methods are highly dependent on the actual magnitude of the effect. Fabian Theis is a professor for Mathematical Modelling of Biological Systems at the TUM and the director of the Institute of Computational Biology at the Helmholtz Zentrum. His team has developed a new measure called kBET which quantifies differences between experiments and therefore facilitates the comparison of different correction results. The findings were presented in a publication in Nature Methods.
An algorithm that detects dropout events
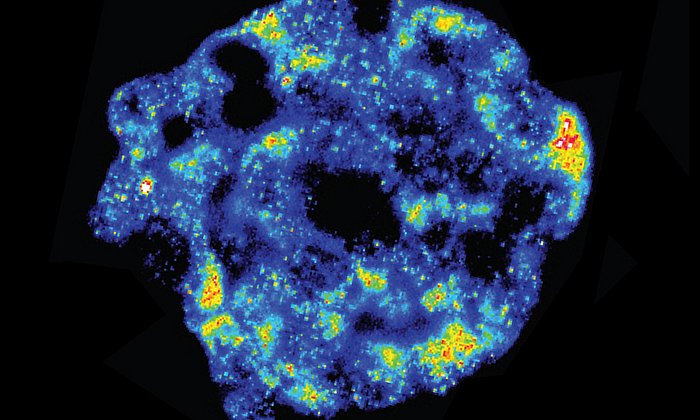
One other challenge for single-cell sequencing are dropout events. "Let’s say we sequence a cell and observe that a particular gene in the cell does not emit any signal at all. The underlying cause of this can be biological or technical in nature: either the gene is not being read by the sequencer because it’s simply not expressed, or it could not be detected for technical reasons", says Fabian Theis.
Whether a dropout event occurred due to a biological or technical failure can now be determined by an algorithm which Theis' group has developed. The software presented in Nature Communications is based on a new probability model and compares the original with the reconstructed data. "We're not developing software to smooth out results", says Theis. "Our chief goal is to identify and correct errors. We’re able to share these data, which are as accurate as possible, with our colleagues worldwide and compare our results with theirs." The reliability and comparability of the data are of paramount importance if they are to be integrated to major projects like the Human Cell Atlas. "Our new algorithm is one of the first in the area of single-cell genomics to be based on neural networks and is the fastest in this field so far", says Theis.
Publications:
Büttner, M.; Theis F. et al. (2019): A test metric for assessing single-cell RNA-seq batch correction. Nature Methods, DOI: 10.1038/s41592-018-0254-1
Eraslan, G.; Simon, L.M.; Theis F. et al. (2019): Single cell RNA-seq denoising using a deep count autoencoder. Nature Communications, DOI: 10.1038/s41467-018-07931-2
ContaCt:
Prof. Dr. Fabian Theis
Technichal University of Munich
Department of Mathematics
Chair of Mathematical Modelling of Biological Systems
Tel.: +49 (0)89 3187 2211
fabian.theis@mytum.de
Technical University of Munich
Corporate Communications Center
- Corporate Communications Center
- lisa.pietrzyk@tum.de
- presse@tum.de
- Teamwebsite