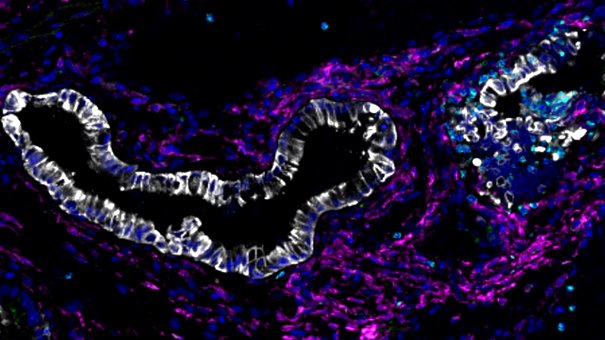
Why identical mutations cause different types of cancer
Cancer development is influenced by tissue type

There have been no major improvements in the treatment of pancreatic and biliary tract cancer in the last decades and no effective targeted therapies are available to date. “The situation for patients with pancreatic and extrahepatic bile duct cancer is still very depressing with approximately only 10% of patients surviving five years,” says Dieter Saur, Professor for Translational Cancer Research at the Technical University of Munich (TUM). Saur is one of the researchers at the German Cancer Consortium (DKTK) partner site at TUM’s university hospital Klinikum rechts der Isar. Joining forces with other researchers at TUM, DKTK, and the University Medical Center Göttingen, Saur has looked into how these forms of cancer develop.
“To discover novel therapeutic strategies that improve prognosis of these patients, it is essential to understand the fundamental genetic networks and interactions that drive these tumors in a tissue-specific fashion. This will allow highly precise molecular interventions in future.”
Unexpected results
The research team looked at the development of biliary tract and pancreatic cancer in mice, replacing the normal “oncogenes” PIK3CA and KRAS with a version containing a mutation identical with that in human cancers. Expression of these oncogenes in the common precursor cells of the extrahepatic bile duct and the pancreas led to very different outcomes. Mice with the mutated PI3K gene developed mostly biliary tract cancer, mice with the mutated KRAS gene instead developed exclusively pancreatic cancer.
This was unexpected because both genes are mutated in both human cancer types. Subsequent analyses discovered the fundamental genetic processes underlying the differential sensitivity of the different tissue types towards oncogenic transformation.
Molecular processes uncovered
“Our studies in mice revealed how genes co-operate to cause cancer in different organs. We identified main players, the order in which they occur during tumor progression, and the molecular processes how they turn normal cells into threatening cancers. Such processes are potential targets for new treatments,” says Chiara Falcomatà the first author of the new publication.
In the mice, the team uncovered a stepwise process of genetic alterations, which drive the development of these cancer types. Some cooperating genetic events overactivate the PI3K signaling pathway, making them cancerous. Others disrupt regulators proteins, inactivating their ability to suppress cancer progression.
Signaling networks susceptible for cancer development
“Understanding the genetic interactions in different cancer types will guide more precise therapeutic decision making in the future” says Günter Schneider, Professor for Translational Cancer Research at the University Medical Center Göttingen.
“What we showed is that the function of an oncogene is different depending on the tissue type and what other genes are altered,” says Roland Rad, Professor for Molecular Oncology and Functional Genomics at TUM and a DKTK researcher. “These oncogenes need to hijack the intrinsic signaling network of a specific tissue to allow cancer development. Interestingly, such networks exist only in specific tissue types making them susceptible for cancer development.”
“No single gene can predict therapeutic response”
These findings have important implications for therapeutic interventions. “The concept that multiple tissue-specific genetic interactions drive cancer progression demonstrates that no single gene can predict responsiveness of a cancer to a particular therapy,” says Saur. “In future, it is key to mechanistically understand the tissue specific determinants of therapeutic response and resistance to get precision medicine to the next level.”
Several of the authors including Dieter Saur and Roland Rad are based at TranslaTUM, TUM’s Center for Translational Cancer Research. In this interdisciplinary research institute, doctors work with colleagues from the fields of natural sciences and engineering on research into causes, diagnostics and potential treatments of cancerous diseases.
C. Falcomatà, S. Bärthel, A. Ulrich, S. Diersch, C. Veltkamp, L. Rad, F. Boniolo, M. Solar, K. Steiger, B. Seidler, M. Zukowska, J. Madej, M. Wang, R. Öllinger, R. Maresch, M. Barenboim, S. Eser, M. Tschurtschenthaler, A. Mehrabi, S. Roessler, B. Goeppert, A. Kind, A. Schnieke, M. S. Robles, A. Bradley, R. M. Schmid, M. Schmidt-Supprian, M. Reichert, W. Weichert, O. J. Sansom, J. P. Morton, R. Rad, G. Schneider, D. Saur. "Genetic screens identify a context-specific PI3K/p27Kip1 node driving extrahepatic biliary cancer". Cancer Discovery 2021, DOI: 10.1158/2159-8290.CD-21-0209
- TranslaTUM website
- DKTK is a consortium centered around the German Cancer Research Center (DKFZ) in Heidelberg, which has long-term collaborative partnerships with specialist oncological centers at universities across Germany.
Technical University of Munich
Corporate Communications Center
- Paul Hellmich
- paul.hellmich@tum.de
- presse@tum.de
- Teamwebsite
Contacts to this article:
Prof. Dr. Dieter Saur
Technical Univerity of Munich (TUM)
Professor of Translational Tumor Research
dieter.saur@tum.de