New mechanism for inhibition of the immunoproteasome revealed
Spot on against autoimmune diseases and chronic inflammations
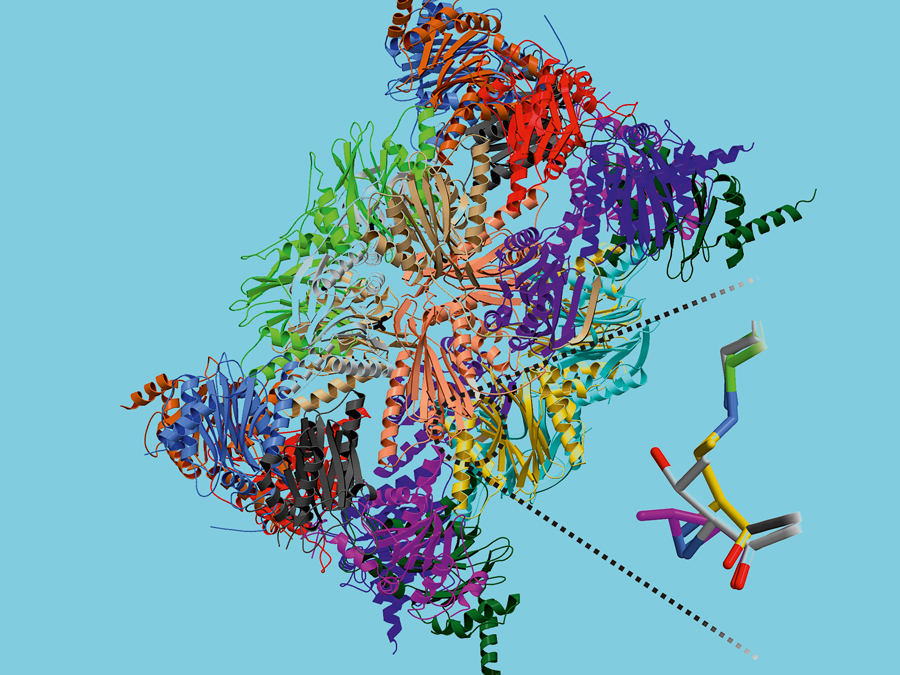
The immune system functions as the body’s police force, protecting it from intruders like bacteria and viruses. However, in order to ascertain what is happening in the cell it requires information on the foreign invaders. This task is assumed by so-called immunoproteasomes. These are cylindrical protein complexes that break down the protein structures of the intruders into fragments that can be used by the defense system.
"In autoimmune disorders like rheumatism, type 1 diabetes and multiple sclerosis or severe inflammations a significantly increased immunoproteasome concentration can be measured in the cells," explains Prof. Michael Groll at the TUM Department of Biochemistry. "The deactivation of this degradation machinery suppresses the regeneration of immune signaling molecules, which, in turn, prevents an excessive immune reaction."
Subtle but significant differences
For some time now, scientists have been on the lookout for new active substances that block immunoproteasomes in a targeted manner without inhibiting the constitutive proteasomes also present in cells. Constitutive proteasomes break down defective or no longer required proteins and are thus responsible for cellular recycling. Notably cell death occurs, when both the constitutive proteasomes and the immunoproteasomes are inactivated.
In early 2012 the research team led by Groll fulfilled a prerequisite for designing specific active substances: They solved the crystal structure of the immunoproteasome, allowing them to spot the subtle but significant differences between the otherwise nearly identical structures.
Special mode of action
The potential drug that the researchers developed is based on the epoxyketon ONX 0914, an immunoproteasome inhibitor that is undergoing clinical trials. The researchers replaced the epoxyketon with a sulfonyflouride group and modified its positioning on the inhibitor. The result was a new compound that selectively inhibits the immunoproteasome without influencing the constitutive proteasome.
First author Christian Dubiella explains what makes the discovered mechanism so special: "Normally inhibitors clog up the active center of the enzyme and thereby disable its functionality. The substance synthesized by us, however, attaches to its target, causing the active center to destroy itself, and then gets detached after successful inactivation." Especially the insights into the atomic mechanisms that were uncovered using X-ray structure analysis open the door to the custom-tailored development of immunoprotease inhibitors. This may pave the road for a future generation of medications.
About the project
The research was done in collaboration with working groups led by Prof. Stephan Sieber at the Department of Organic Chemistry II and Prof. Achim Krüger at the Institute of Experimental Oncology and Therapy Research at TUM, as well as Prof. Robert Liskamp at the University of Glasgow. The work was funded by the German Research Foundation DFG (SFB 1035/A2 & DFG GR 1861/10-1), as well as the Excellence Cluster Center for Integrated Protein Science Munich (CIPSM). The X-ray diffraction measurements were conducted at the PXI Beamline of the Paul Scherrer Institute (Villingen, Switzerland).
Original publication:
Selective Inhibition of the Immunoproteasome by Ligand-Induced Crosslinking of the Active Site, Christian Dubiella, Haissi Cui, Malte Gersch, Arwin J. Brouwer,Stephan A. Sieber, Achim Krüger, Rob M. J. Liskamp, Michael Groll, Angewandte Chemie Int. Ed., DOI: 10.1002/anie.201406964
Download high-resolution picture
Contact:
Prof. Dr. Michael Groll
Technische Universität München
Department of Biochemistry
+49 89 289 13361
michael.groll@tum.de
http://www.biochemie.ch.tum.de/