New mechanism in gene regulation revealed
Doubled productivity
The number of genes in humans, mice and fruit flies is almost identical, of around 20.000, and thus cannot explain the phenotypic differences between these organisms. Thus, not only the number of genes but also their regulation plays a critical role for the evolution of life.
Upon transcription of the DNA, mRNA is produced as an initial step, which subsequently functions as template for the synthesis of proteins. Protein synthesis is regulated by the binding of regulatory proteins to the mRNA. How these proteins specifically recognize the RNA has been poorly understood to date.
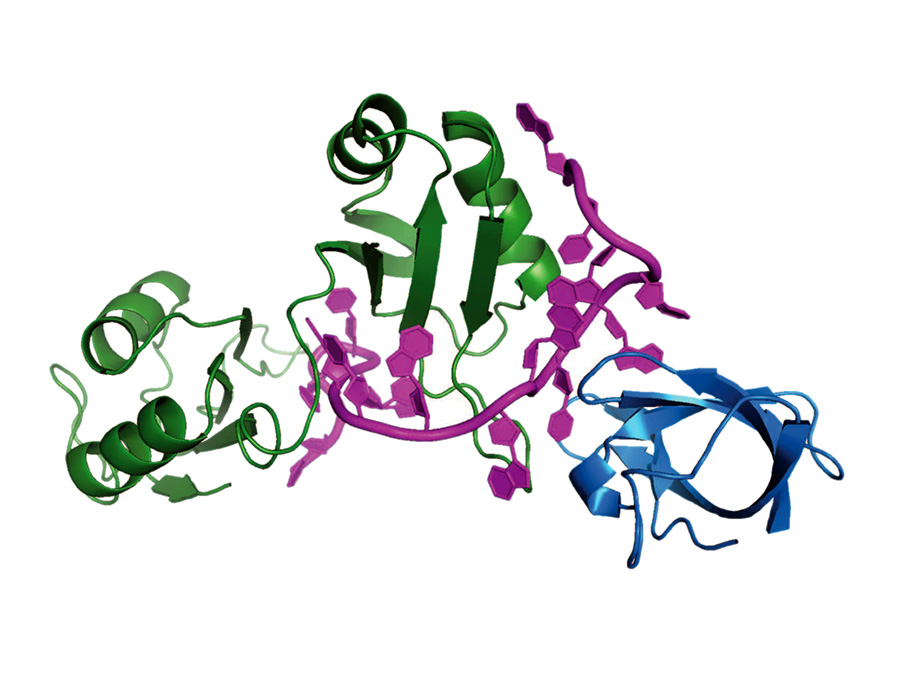
An international team, led by Helmholtz Zentrum München (HMGU) and Technische Universität München (TUM), has now determined the three-dimensional structure of such a regulatory protein-RNA-complex. The scientists studied the specific complex formed between the regulatory proteins Sxl (Sex-lethal) and Unr (Upstream-of-N-Ras) with mRNA in the fruit fly model
This protein complex is essential to ensure that the expression of genes located on the two female (XX) X-chromosomes equals that of genes located on the single male (XY) X-chromosome. This guarantees that comparable protein concentrations are achieved in both sexes, which is essential for viability of the fruit flies.
A paradigm for gene regulation
The three-dimensional structure now reveals how multiple proteins collaborate for highly specific recognition of the mRNA. “Our results represent a paradigm for the regulation of various essential cellular processes at the level of mRNA”, explains Professor Michael Sattler, Professor for Biomolecular NMR Spectroscopy at TUM and director of the Institute of Structural Biology at HMGU.
The specific recognition of mRNA is achieved by cooperation of several RNA-binding proteins, even though these proteins individually exhibit poor binding affinity and are involved in distinct processes in the cell. By combining multiple proteins, the number and variety of biological processes that can be regulated by a relatively small number of regulatory RNA binding proteins, is greatly expanded.
The authors expect that this principle represents an essential and widespread mechanism of gene regulation in higher organisms, where mutation or misregulation of homologous proteins has been implicated in disease.
The research was conducted by a group of scientists around Dr. Janosch Hennig, Dr. Grzegorz Popowicz and Professor Dr. Michael Sattler together with the group of Dr. Fátima Gebauer at the Centre for Genomic Regulation (Barcelona, Spain) and colleagues at the Institut Biologie Structurale (Grenoble, France). X-ray crystallography and Nuclear Magnetic Resonance spectroscopy experiments were performed at the Institute of Structural Biology of HMGU and the Bavarian NMR Centre (TUM and HMGU). Small angle scattering was done at the Institut Laue-Langevin and the Institut Biologie Structurale (Grenoble, France).
Publication:
Hennig, J. et al., Structural basis for the assembly of the SXL-UNR translation regulatory complex, Nature, DOI: 10.1038/nature13693
Contact:
Prof. Dr. Michael Sattler
Technische Universität München
Chair for Biomolecular NMR-Spectroscopy
Lichtenbergstr. 2, 85748 Garching, Germany
Tel.: +49 89 289 13867 – E-Mail – Internet