Chaperones detect immature signaling molecules in the immune system
Quality control in immune communication
The body's defenses systems have to react quickly whenever pathogens enter the organism. Intruders are identified by white blood cells which pass on the information to other immune cells. Information is transmitted via secreted signaling proteins, the interleukins, which dock onto the matching receptors on the recipient cells and for example make the target cells divide and release antibodies.
Quality control holds back immature signaling molecules
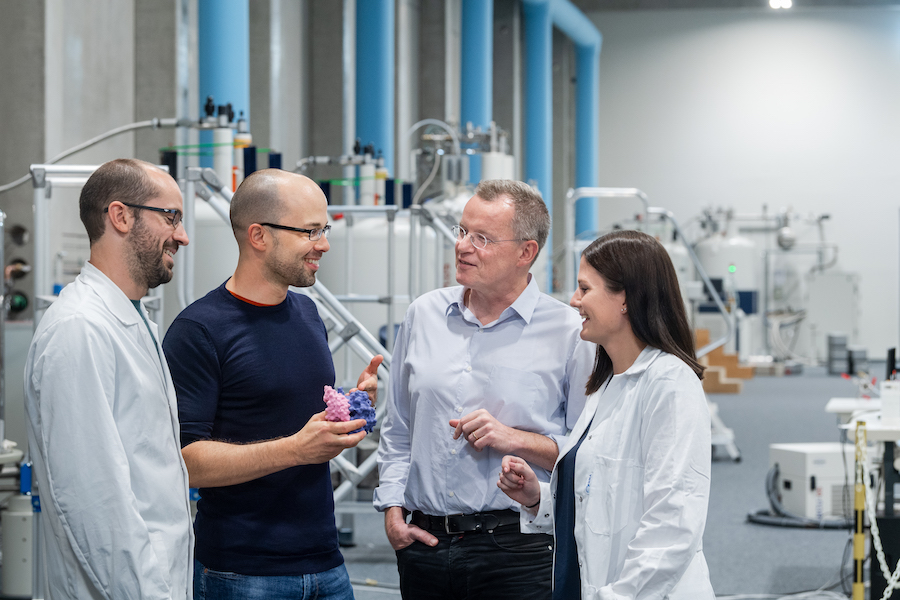
Researchers from TUM, the Helmholtz Zentrum München and Stanford University have, by studying interleukin 23, been able to show how cells ensure that the interleukin signalling proteins are built correctly. "Intensive research is currently devoted to Interleukin 23, not only because of its central role in the defense against pathogens, but also because it can trigger autoimmune diseases," says Matthias Feige, Professor for Cellular Protein Biochemistry at TUM and head of the research project.
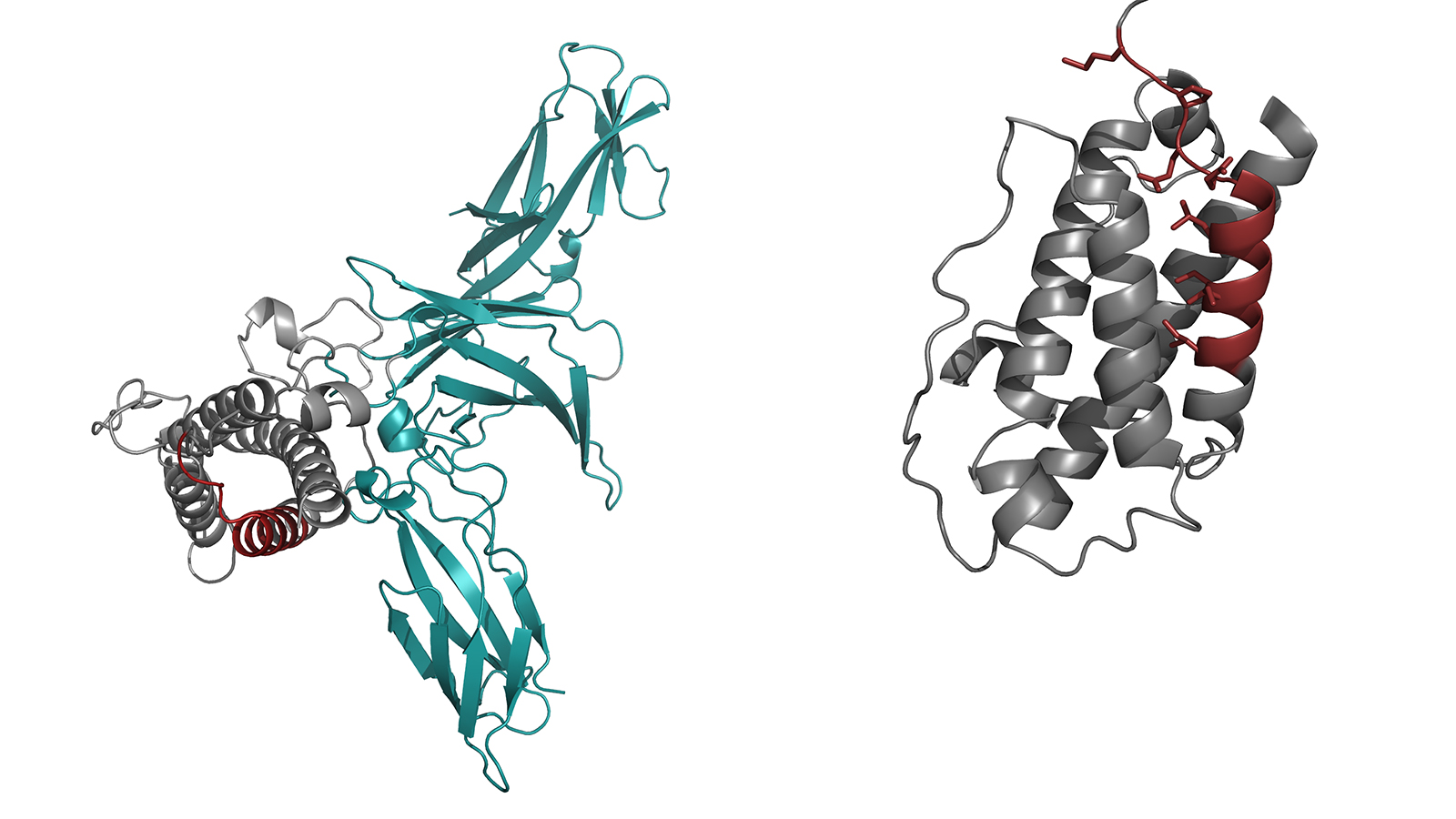
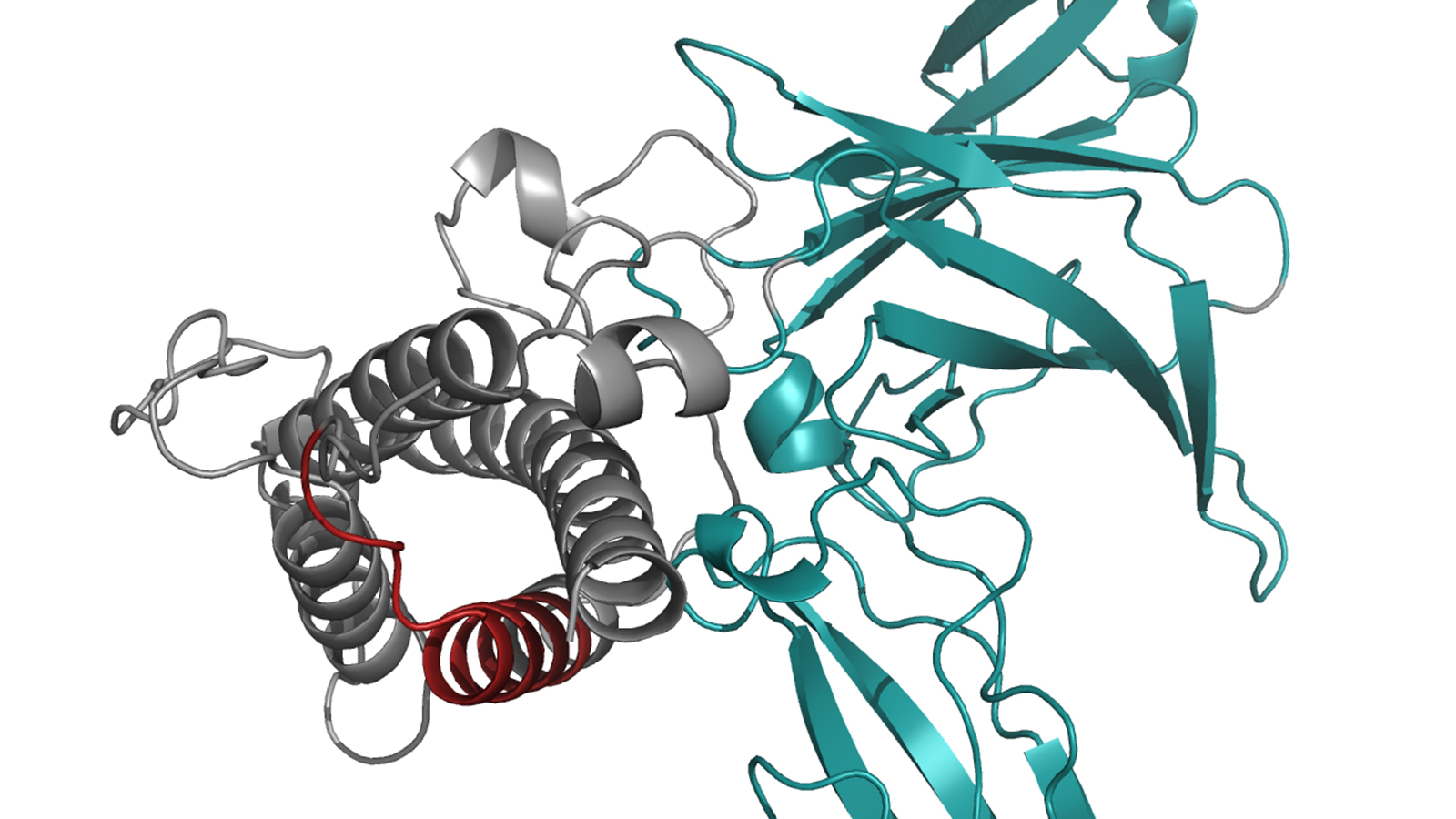
Interleukin 23 is composed of two proteins, which have to combine in the cell to form an active complex in order to be able to trigger the desired signals. As the scientists have demonstrated in their study, molecules referred to as chaperones retain one part of the interleukin known as IL23-alpha in the cell until it has been incorporated into the complete complex. This way the cell makes sure that it does not secrete any unpaired IL23-alpha and thus controls the biosynthesis of this important interleukin and accordingly of the messages it sends. Chaperones are molecular protein machines that ensure that other proteins are built correctly.
"We were able to show that unbound IL23-alpha has chemical bonds which are prone to interaction with chaperones," Feige explains. In the completed interleukin 23 these bonds are closed, so that the chaperon no longer is able to interact and hence the complete molecule can leave the cell.
Targeted interventions in immune cell communication
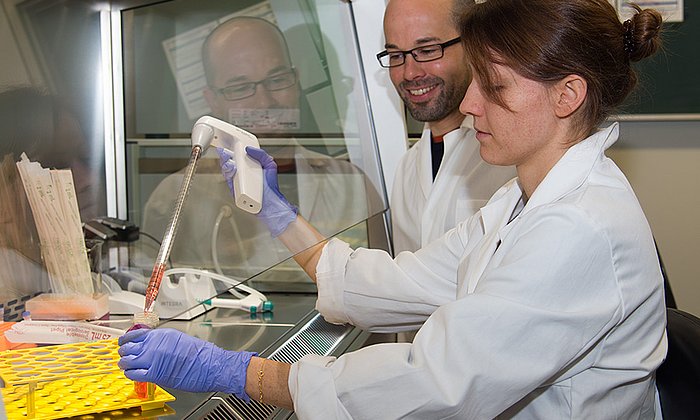
Since normally isolated IL23-alpha is not present outside of the cell, it was not clear whether it could influence the immune system by itself. The researchers were able to test this with a slightly modified version of the molecule created in the laboratory, which was based on computer-aided design. In this new molecule variant, the bonds which could have connected to the chaperone were closed.
"The modified molecules can leave the cell freely," says Susanne Meier, first author of the study. "They then dock to the same receptors as the complete interleukin 23 and trigger a similar but weaker reaction." Accordingly, IL23-alpha can be made a functional signalling protein by molecular engineering, which allows it to bypass the cell's quality control systems.
"It is possible that the engineered IL23-alpha can interact with even further receptors in immune cells and influence them in an as yet unknown manner," Feige says. "That is one of the next questions we will investigate." The results may serve as the basis for future drugs that use engineered interleukins to modulated the immune system in a desired manner.
S. Meier, S. Bohnacker, C. J. Klose, A. Lopez, C. A. Choe, Ph. W.N. Schmid, N. Bloemeke, F. Rührnößl, M. Haslbeck, J. Esser-von Bieren, M. Sattler, Po-Ssu Huang & M. J. Feige. “The molecular basis of chaperone-mediated interleukin 23 assembly control”. Nature Communications 10, 4121 (2019); DOI: 10.1038/s41467-019-12006-x
Matthias Feige’s professorship is funded by the German Research Foundation's Excellence Initiative and by the Seventh Framework Program of the European Union under Grant Agreement 291763, through the TUM Institute of Advanced Study and the Marie Curie COFUND program. The research was conducted as part of the Collaborative Research Center (SFB) 1035 "Control of Protein Function by Conformational Switching", projects A03, B11 and Z1.
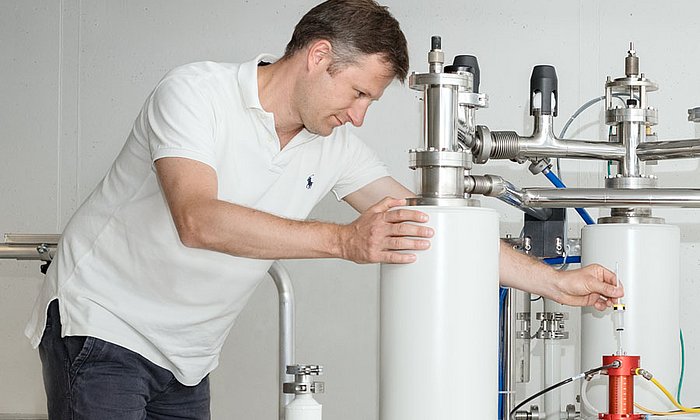
Researchers from the Center of Allergy & Environment (ZAUM) played a central role in the project. Structural investigations of the proteins were conducted at the Bavarian NMR Center. Both facilities are joint institutions of TUM and the German Research Center for Environmental Health Helmholtz Zentrum München.
The image of the protein structure is based on results published in the article:
Lupardus, P. J. & Garcia, K. C. "The structure of interleukin-23 reveals the molecular basis of p40 subunit sharing with interleukin-12“. J. Mol. Biol. 382, 931–941 (2008) doi: 10.1016/j.jmb.2008.07.051
The data is available from the Protein Data Base under the ID 3DUH.
High resolution images:
mediatum.ub.tum.de/1519865
Contacts to this article:
Prof. Dr. Matthias Feige
Technical University of Munich
Cellular Protein Biochemistry
Tel.:+49 89 289 13667
E-Mail: matthias.feige(at)tum.de